Po-Yu Kao, Ya-Chu Yang, Wei-Yin Chiang, Jen-Yueh Hsiao, Yudong Cao, Alex Aliper, Feng Ren, Alan Aspuru-Guzik, Alex Zhavoronkov, Min-Hsiu Hsieh, Yen-Chu Lin
Journal of Chemical Information and Modeling (JCIM)
Abstract
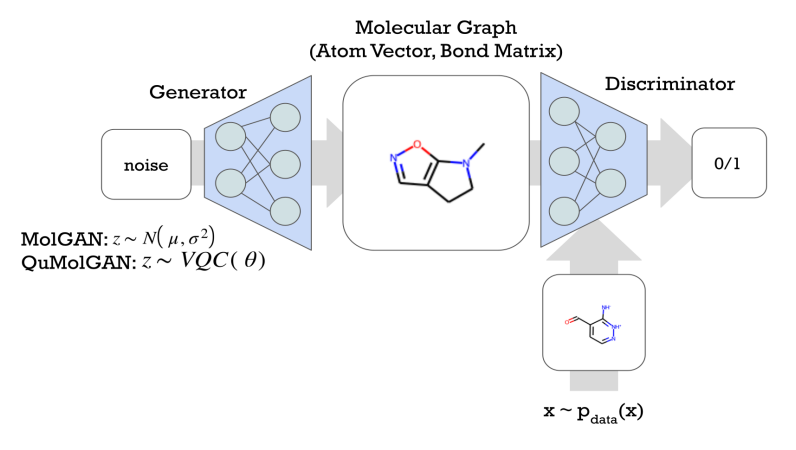
De novo drug design with desired biological activities is crucial for developing novel therapeutics for patients. The drug development process is time and resource-consuming, and it has a low probability of success. Recent advances in machine learning and deep learning technology have reduced the time and cost of the discovery process and therefore, improved pharmaceutical research and development. In this paper, we explore the combination of two rapidly-developing fields with lead candidate discovery in the drug development process. First, Artificial intelligence has already been demonstrated to successfully accelerate conventional drug design approaches. Second, quantum computing has demonstrated promising potential in different applications, such as quantum chemistry, combinatorial optimizations, and machine learning. This manuscript explores hybrid quantum-classical generative adversarial networks (GAN) for small molecule discovery. We substituted each element of GAN with a variational quantum circuit (VQC) and demonstrated the quantum advantages in the small drug discovery. Utilizing a VQC in the noise generator of a GAN to generate small molecules achieves better physicochemical properties and performance in the goal-directed benchmark than the classical counterpart. Moreover, we demonstrate the potential of a VQC with only tens of learnable parameters in the generator of GAN to generate small molecules. We also demonstrate the quantum advantage of a VQC in the discriminator of GAN. In this hybrid model, the number of learnable parameters is significantly less than the classical ones, and it can still generate valid molecules. The hybrid model with only tens of training parameters in the quantum discriminator outperforms the MLP-based one in terms of both generated molecule properties and the achieved KL divergence.
[Link]